With one-third of all food produced globally lost or wasted each year, developing the most effective preservation strategy has never been more important. Yet the challenge goes far beyond simply adding a “one-size-fits-all” preservative. Have you ever wondered why your lactate-based preservative may work very well in cooked ham but is far less effective in a plant-based product? An important reason could be that the main spoilage microorganisms are not the same across different food products.
Every food matrix has its own unique microbiological profile, which can be determined by many different factors such as origin, processing methods, storage conditions, physicochemical properties... These parameters determine which microorganisms will thrive or not. Therefore, to identify potential contamination risks and determine appropriate preservation methods, manufacturers must have a thorough understanding of the microorganisms present in a product. This is where metagenomics provides a powerful solution. Such an approach aligns with the conclusions of the March 2026 FAO/WHO joint meeting on the use of omics tools, which highlighted metagenomics as a key technology for improving food safety assessments, microbial risk characterization and evidence-based decision-making.
Seeing microorganisms in action
Metagenomics is a DNA analysis technique used to identify microorganisms present in a (food) sample. It provides a complete picture of the microbiota of the sample. In the context of food preservation, the strength of metagenomics lies in its ability to track the evolution of the microbial profile over time. By observing which populations grow and become dominant, it becomes possible to identify the microorganisms that could compromise product quality and safety. This is important because the presence of a microorganism at the beginning of a product’s shelf-life does not automatically imply a risk. Some products such as yoghurt, some cheeses or fermented products have a high microbial concentration at the beginning of their shelf-life but this does not make them unsafe for consumption.
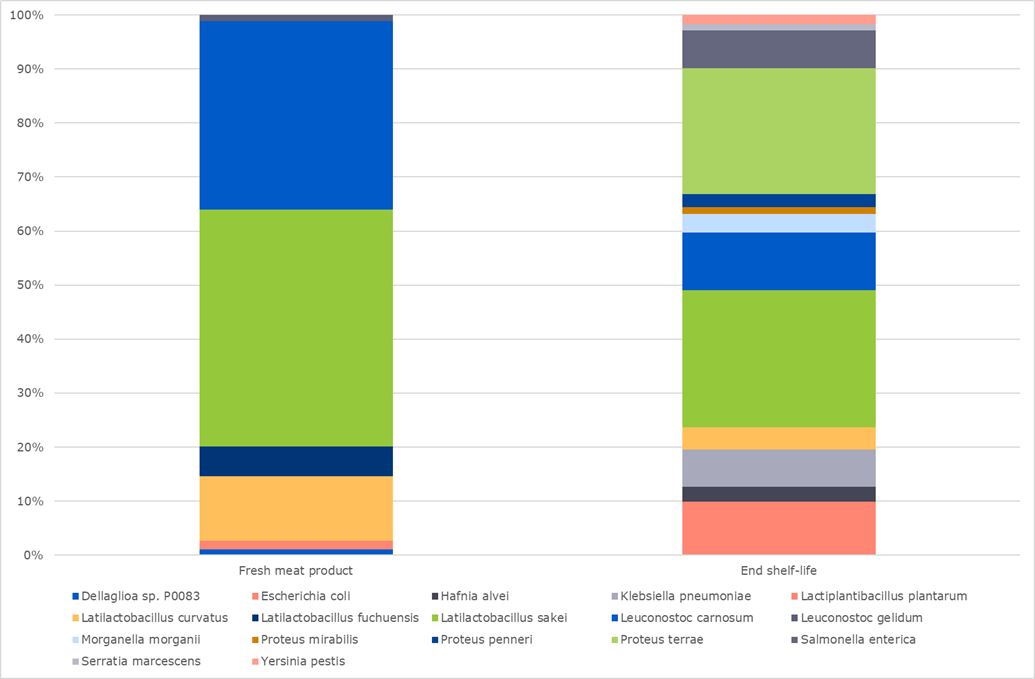
In figure 1, two meat products from the same production batch are compared: one at the beginning of its shelf-life and the other after the end of its shelf-life. In the fresh sample, Latilactobacillus sakei and Leuconostoc carnosum were the predominant species, representing 80 % of the bacterial population. By the end of the shelf-life of the product, these species accounted for only 45 %, indicating that they were not the main bacteria that proliferated during storage.
On the contrary, Proteus terrae, Proteus penneri and Proteus mirabilis were initially present at low levels but grew significantly to represent up to 30 % of the bacterial population towards the end of the shelf-life. This analysis shows that the primary source of degradation of the meat product is mostly due to the growth of Proteus spp. rather than the initially more abundant Latilactobacillus sakei and Leuconostoc carnosum.
Tailored preservation strategies
Knowing exactly which microorganisms are present in a product gives producers a precise understanding of potential risks and preservation needs. Such an insight allows them to select the most effective preservatives, avoiding overuse while ensuring optimal protection and product quality. Furthermore, metagenomic enables a smarter shelf-life optimization strategy, reducing “trial-and-error” adjustments and guesswork during the R&D phase. For instance, if unexpected spoilage appears, such as a white liquid on a product, it could be microbial or result from another cause. Metagenomics can determine whether microorganisms are involved and identify the responsible species.
By combining targeted preservation with a clear view of microbiological risks, producers can lower costs, safety incidents and build consumer trust. This approach will also protect the brand and meet growing demand for safe, transparent, and high-quality food.
In the example in the previous section (figure 1), Proteus spp. were identified as the main source of spoilage. They belong to the Enterobacteriaceae family and are Gram-negative, facultative anaerobic bacteria, meaning they can grow with or without oxygen. Their presence indicates potential hygiene issues in the production environment and highlights risks that could compromise product quality and safety, such as biofilm formation, gas production, or tolerance to environmental stresses. Recognizing these traits allows manufacturers to implement targeted interventions, such as enhanced hygiene protocols, tailored preservatives, and optimized storage conditions, to effectively mitigate spoilage risks and maintain consistent product quality.
By understanding the source of spoilage, the most (cost-)effective preservation strategy can be elaborated using different hurdles. Preventing initial contamination and maintaining strict hygiene in processing areas will be critical. Adding extra hurdles such as refrigerated storage and modified atmosphere packaging will help protect the product throughout the entire shelf-life. Finally, a key hurdle would be the addition of a targeted preservative. Because facultative anaerobic bacteria are the main spoilage organisms in this case, adding (di)acetate is effective. These facultative anaerobic bacteria can switch between energy systems to support growth and maintain internal balance, such as pumping out protons to regulate pH. Adding undissociated acetate disrupts this by entering the cell and lowering its internal pH, forcing the bacteria to expend energy restoring homeostasis. This acid stress interferes with essential metabolic processes, slowing growth and making acetate an effective way to control spoilage and extend shelf-life.
Understanding the boundaries
While metagenomics provides a powerful and comprehensive view of microbial populations, it is important to recognize the limitations of this approach and the complementary role of classical methods.
Classical enumeration methods measure the growth of microorganisms on selective media. They only allow for the detection of species capable of developing under laboratory conditions. In addition to being time-consuming, such methods can miss certain microorganisms called “viable but non-culturable”. Such microorganisms are still alive but unable to grow on standard culture media. This phenomenon often occurs when microorganisms are subjected to stresses such as cold, salt or acidity.
By contrast, metagenomics analyzes all the DNA present in a given sample. Thus offering a much broader view of microbial diversity, including species that are difficult to culture. However, this approach primarily provides relative abundances of species rather than absolute quantities. Thus, while a species might appear to increase by 30% in relative terms, the overall population may have changed very little, limiting the biological interpretation. Metagenomics also does not provide information on microbial activity and therefore does not distinguish between living, dormant or dead cells. It is also more expensive and requires greater expertise.
When combined with classical enumeration, metagenomics becomes a much stronger tool because changes in DNA profiles can be linked to actual microbial growth and activity. By comparing DNA data with microbial growth on selective media, it is possible to determine whether microorganisms are actively growing, remaining dormant or already dead, and this combined approach greatly reduces the uncertainty of using DNA data alone while providing a clearer picture of which microorganisms are truly active.
This is why, at Galactic, we have implemented metagenomics in our laboratory to complement our current classical enumeration techniques. By combining these two approaches, we gain a more complete and reliable understanding of the microbial populations present and the potential associated risks. This insight enables us to guide customers in selecting the most effective preservative.

Smarter shelf-life, safer food
Effective food preservation starts with understanding the microorganisms that are present and growing in each product. Metagenomics, combined with classical enumeration methods, provide a complete and precise picture of microbial risks, helping producers to choose the most targeted preservation strategies. This approach not only leads to extended shelf-life and reduced spoilage but also ensures safety, lowers costs and builds consumer trust. In a market where transparency and quality increasingly matter, data-driven and tailor-made preservation is the key to protecting both products and brand reputation.
By Héloïse de Wilde, Food Applications Engineer at Galactic

Still have questions?
Can’t find the answer you’re looking for?
Contact us for expert advice and support!


